Spatial Omics: Nanostring GeoMX & CosMX
Determine RNA or protein profiles based on morphology from small cell populations to within single cells
Digital Spatial Profiling: GeoMX
Current techniques offer a trade-off between spatial data and in-depth expression data. Bulk sequencing, for instance, generates in-depth data, but often it tells us little about where these cells were situated in the original tissue. Techniques like single molecule FISH and immunohistochemistry offer us spatial data, but only have a limited amount of targets. USEQ offers a new technique called GeoMX (Nanostring), which combines the spatial data with RNA or protein expression profiles. GeoMX opens up a world of research possibilities. You provide the tissue of your choice and select regions that interest you. This creates a wide range of applications from creating separate profiles for tumor and tumor microenvironment, to finding biomarkers for neurodegenerative diseases, to the effects of COVID infection on gene expression.
FFPE-preserved tissues
Many RNA-based techniques need fresh-frozen tissue. One of the unique strengths of GeoMX is the input of a multitude of types of tissue preservations, including FFPEpreserved tissue. Multi-omics technique Because it is possible to obtain both protein profiles and RNA profiles with GeoMX, GeoMX is considered a multiomics technique. Its strength lies in being able to create datasets of RNA on one slide, and protein datasets on an adjacent slide. These two could then be compared to each other, in order to gain conclusions which would not be possible with only proteomics or transcriptomics.
Process
We stain tissue slides with RNA/protein detection reagents and morphology markers, such as for epithelium, immune cells and nuclei. We are open to custom morphology markers. The tissue is then processed by the Digital Spatial Profiler, creating a high-quality image on a single cell visibility level. We then select regions of interest, in which we can select different cell types based on morphology staining.
These areas are then illuminated, cleaving the RNA/protein reagents only in the illuminated areas. This enables a highly specific region to be selected. These area’s of illumination are collected in different wells. We then sequence this data from the selected regions and generate RNA or protein expression profiles per region or cell type. 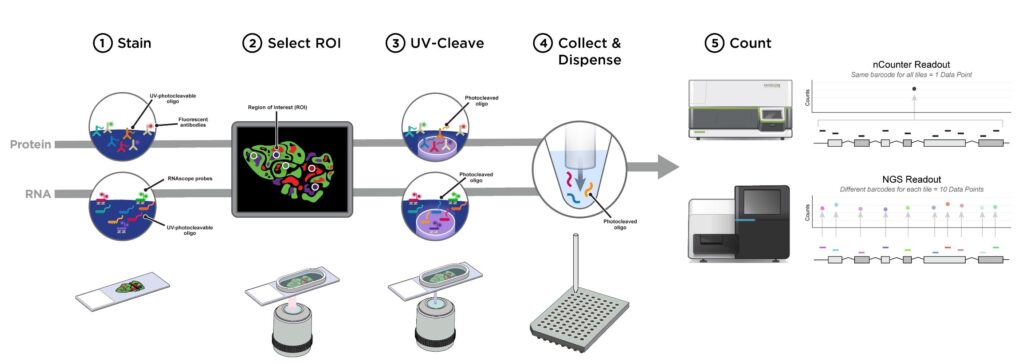
CosMX: Single Cell, up to sub-cellular locations
The latest generation spatial omics by Nanostring is CosMX. This takes the combination of spatial information and detailed RNA profiles a step further, because it can do this on a single-cell level. It does so by generating highly detailed cell segmentation and creating a 1000-plex RNA transcripts image, which enables us to see thousands of RNA’s on a single cell level. The CosMX machine is available in USEQ for running both RNA and protein projects.
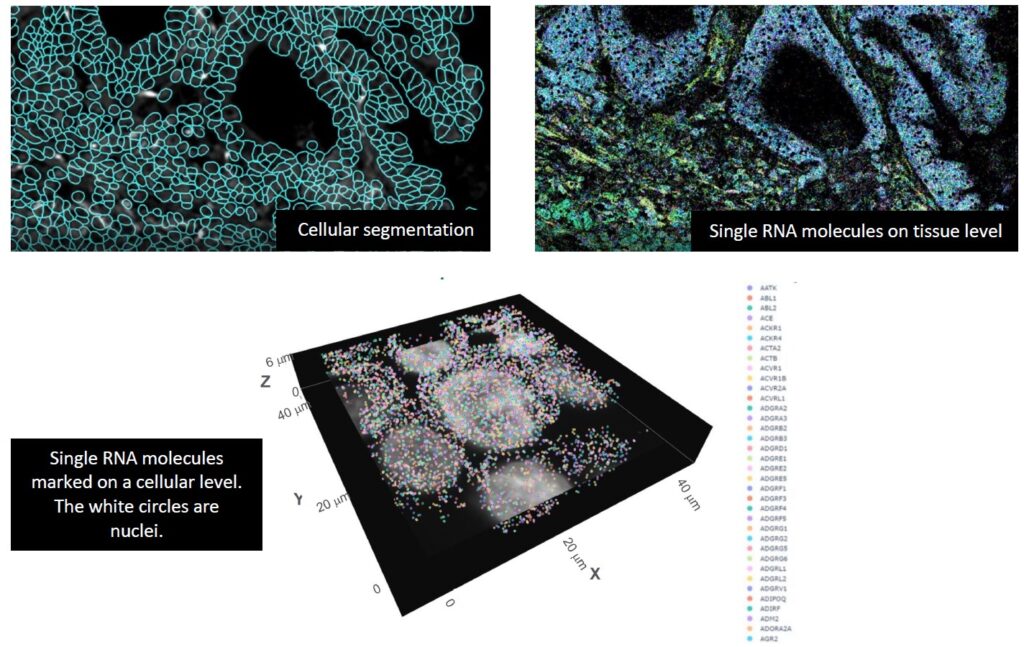
The CosMx™ SMI and decoder probes are not offered and/or delivered to the Federal Republic of Germany for use in the Federal Republic of Germany for the detection of cellular RNA, messenger RNA, microRNA, ribosomal RNA and any combinations thereof in a method used in fluorescence in situ hybridization for detecting a plurality of analytes in a sample without the consent of the President and Fellows of Harvard College (Harvard Corporation) as owner of the German part of EP 2 794 928 B1. The use for the detection of cellular RNA, messenger RNA, microRNA, ribosomal RNA and any combinations thereof is prohibited without the consent of the of the President and Fellows of Harvard College (Harvard Corporation).
Sample Submission
For sample submission please follow the procedure described here.
Pricing
| Service | Price per | Price (excl VAT) |
|---|---|---|
| GeoMX | Run | € TBD |
| CosMX | Run | € TBD |
Service fees include costs for chemicals, operator time and troubleshooting.
Consultancy Service
For expert advice on your NGS-based research and the possibilities USEQ offers to get the most out of your samples: request a meeting at our consulting hour, each Friday at 1:30pm. Experimental design, technology, platform and library preparation choice, standard bioinformatics analysis, results and follow-up experiments are discussed. The facility manager ensures to bring the right experts from any of the participating organisations to the table.
You can request a meeting at useq@umcutrecht.nl.
